Overview of GDI data operations
- Version: 0.5
- Status: DRAFT
- Created: 2026-01-20
- Updated: 2026-01-20
- To be consulted:
- Kersti Jääger (kersti.jaager@ut.ee)
- Erik Jaaniso (erik.jaaniso@ut.ee)
Introduction
As part of the European GDI project, Estonian GDI (GDI-EE) team at Institute of Computer Science, University of Tartu has developed an IT infrastructure that exposes national genomic dataset information to the central European User Portal for harmonised search of genomic data. The team has developed data pipelines that can prepare and ingest data and its associated metadata in a structured format and convert it into accessible yet protected data tables within its secure storage system. A graphical user-friendly web interface (GUI) was designed for data-providers to easily submit and manage their datasets. Dataset management includes the possibility to change dataset status and thereby control dataset visibility in the User Portal. The UI also enables data-provider organisations to have control over their accounts and associated permissions, a roles management feature not detailed in this piece of writing. Hereby, descriptions about data operations in the GDI-EE system are delivered to the node-holder organisation.
This overview describes processes that cover GDI-EE dataset lifecycle from data ingestion to data access via User Portal. Dataset transfer to SAPU, a sensitive data analysis platform at UT HPC (SAPU), and subsequent analyses in SAPU are only briefly listed for system completeness purposes, yet these processes will be entirely managed by UT HPC and will involve contractual operations established by the involved parties. Together with initial risk mapping (Risk Mapping), this information is aimed to support the node-holder organisation in understanding the logic behind GDI-EE data flow, identifying potential data privacy risks, and ensuring compliance with data protection regulations and organisational policies.
Dataset processing lifecycle
GDI-EE infrastructure architecture facilitates the processing of datasets along sequential steps in the dataset lifecycle. The data pipelines, preparation and managing steps in GDI-EE are together designed to support:
- The use of Variant Call Format (VCF) as an input data which will be converted to a variant discovery file that can be queried from via User Portal.
- The ingestion of Metadata in YAML format that is structured according to HealthDCAT-AP standard to enable dataset discovery via User Portal.
- The dataset registration and state management by data-providers in (local node portal).
For a structured description of the data operations, we first divided them into six core steps: 1) Dataset package preparation, 2) Dataset ingestion, 3) Dataset management, 4) Data search and access, 5) Data transfer to SAPU, 6) Data analysis in SAPU (Diagram 1), each detailed below. However, while handing over GDI solution to the future node-holder (2026): only operations 1-3 have been fully specified by GDI-EE; operation 4 being highly dependent on User Portal developments still underway; and operations 5-6 being specified by node-holder and involved parties after the GDI project.
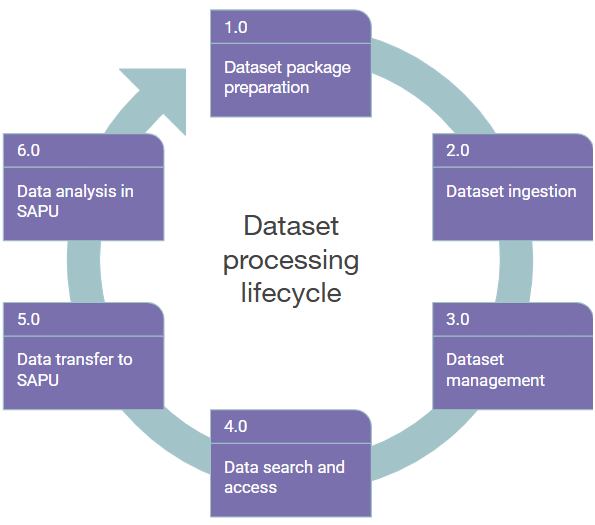
Diagram 1. Dataset processing lifecycle of GDI-EE starts with creating a dataset package by data-provider (1) that will be ingested and registered in the GDI system (2). Dataset management (3) includes operations that data-providers can perform with registered datasets. Dataset properties are exposed to User Portal where a researcher can find and request access to data (4). Data transfer to and analysis in SAPU (5, 6) remain to be coordinated by data-provider and node-holder organisation to support secure environment setup and obtaining of research results.
Dataset package preparation (1.0)
Genomic variant data and associated metadata for the dataset and samples (optional) needs to be packaged by the data-provider before it can be entered into the GDI system. The packaging involves basic sanity checks for the content and match between the provided files (VCF, package.yaml, samples.csv) and conversion of original files into Parquet files and validating metadata to manifest file (manifest.yaml). The samples file should be omitted in case of summary-level datasets. See the list of differences between summary- and individual-level datasets in terms of content and sensitivity in Table 1. The resulting output is an encrypted dataset package file (.tar.c4gh) that can be deposited in the Shared storage (Minio S3) by the data-provider. See gdi-dataset-tool and compiling-a-dataset-package for technical details.
Dataset ingestion (2.0)
Dataset package can either be uploaded in the local portal or put into data-provider’s S3 storage for dataset ingestion into GDI-EE. The ingestion pipeline performs several processing steps including a) adding a new database record for unregistered dataset, b) validating files content of the package, c) updating database records (see Database tables in Table 2), d) encrypting Parquet files using keys from Vault, and e) saving files in node S3 storage. Upon successful validation, the package files will be removed from the data-provider’s storage and dataset status will be updated to REGISTERED in the local portal. The requirements for the package files to pass validation are described here: manifest file (dataset-manifest), variant files (beacon-variants, beacon-allele-frequencies), samples file (dataset-samples). At this stage, dataset details are only accessible to the data-provider.
Dataset management (3.0)
Each dataset that is registered in the GDI-EE can be tracked and managed by the user via the GUI of the local portal. To control the status of the dataset, the user can choose between REGISTERED -> VISIBLE -> HIDDEN -> ARCHIVED statuses in the dataset lifecycle (datasets). However, to publish a dataset in the User Portal, dataset status must become VISIBLE. As a result, dataset is listed in the local FAIR Data Point catalogue where it is retrieved by the User Portal (fair-dp), and indexed data files are exposed to Beacon search operations. VISIBLE dataset and its files are given persistent GDI-EE identifiers. Local dataset management also allows to make a dataset invisible for the User Portal (HIDDEN) or turn it into ARCHIVED state which removes Beacon files from the storage (keeping only dataset metadata).
Table 1. Differences between summary- and individual-level data discovery services deployed by GDI nodes.
| Summary-level data | Individual-level data | |
|---|---|---|
| EU User Portal | Allele Frequency Browser | Data Catalogue |
| Beacon service | Allele Frequency Beacon | Sensitive (Regular) Beacon |
| Beacon instance | Aggregated Beacon | Subject-level Beacon |
| Access level | Public | Non-public* |
| Data sensitivity | Non-sensitive | Sensitive |
| Samples file in dataset package | No | Yes |
| Use case type | Genome of Europe variant data including pre-computed cohort-based allele frequencies (real) | Variant data supplemented with samples’ phenotypic data (synthetic) |
| Data structure | Specific for GoE project | Aligns with 1+MG HMD |
| Cohort | Population average | Patients |
| Beacon response | Record | Count |
| Authentication | Optional | Required |
| GDI Maximal Attainable Product (October 2026) | Required | Optional |
| Readiness | Production | Proof-of-concept |
*Implemented as Public at the moment using synthetic data (April 2026)
Data search and access (4.0)
EU-level dataset search is described by two distinct use cases that are supported by two Beacons: 1) Aggregated Beacon via User Portal Allele Frequency Browser for querying Genome of Europe allele-frequency data, 2) Subject-level Beacon via User Portal Data Catalogue for querying synthetic variant data. Note that the requests are similar for both Beacons whereas the minimal Beacon API response is either record (1) or count (2). Search filters for dataset are implemented according to HealthDCAT-AP v5 specification, while phenotypic filters are defined by Genome of Europe pilot use case, including pre-defined cohorts (metadata-of-genomic-datasets). Data search is possible for datasets that are exposed in the data catalog.
Due to the unfinalised state of the Genome EDIC, no central access governance procedures have been defined or implemented. Therefore, researchers’ flow from finding an interesting dataset, filing an access application and receiving a permission for data analysis are still completely handled case-by-case via manual processes (see rems–the-permission-service for explanation). Therefore, no authentication (using Life-Science AAI) has been deployed in the User Portal to enable controlled access at this stage of the GDI project. All searchable data is classified as PUBLIC. Although Subject-level Beacon is by its nature designed for sensitive data (Table 1), no controlled-access Beacon requests are supported by current implementation.
Data transfer to SAPU (5.0)
We propose, however, that when a data-user has been granted access to genomic data, a cohort description file (Data cohort manifest) is generated that includes only data files specified in the access permission document. These files (Data cohort files) can be securely transferred from Shared storage to the SAPU data analysis environment. SAPU preparation and management processes are regulated according to service-provider rules.
Data analysis in SAPU (6.0)
Again, we propose that data analysis in GDI-EE takes place in SAPU at University of Tartu. No original data file can be downloaded from SAPU. Analysis results can be exported only upon receiving a permit from a dedicated authoritative organisation as specified in the Genome EDIC.
Data flow diagram
To visualise the connection between the described processes and sub-processes as well as the data stores they interact with, a data flow diagram structure has been used (Diagram 2). The main benefit of a data flow diagram is visualising processes that ‘do’ something with the data, and stores that ‘keep’ data in a certain file or location. This approach helps to visualise operations that are performed with data on top of the GDI-EE architecture.
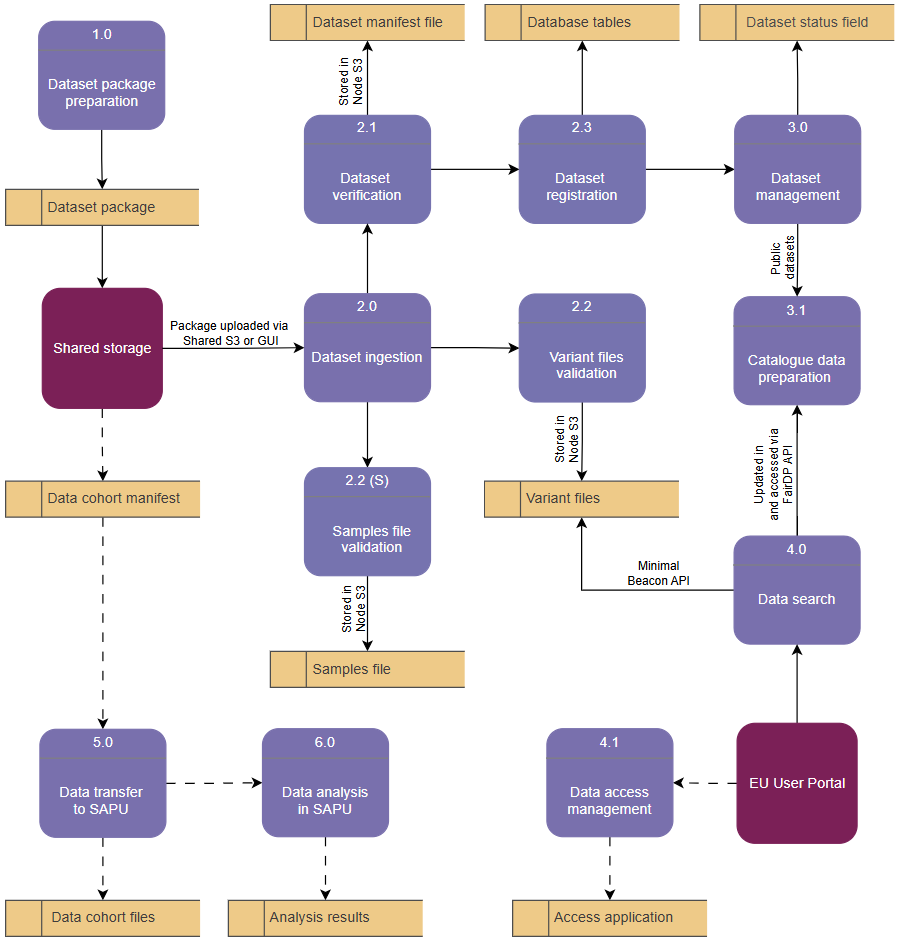
Diagram 2. Data flow diagram showing data stores (yellow) associated with the relevant processes or sub-processes (purple) which have connection with two external entities (bordeaux): Shared storage (data-provider interaction with the system) and EU User Portal (data-user interaction with the system). The (S) for Samples file validation denotes an optional process only applicable to subject-level data used with Sensitive Beacon. Dashed lines indicate processes that have not been fully defined by the GDI project.
Data stores associated with data operations
Data in the node system is imported, processed and saved in diverse formats for use in specific scenarios. The following table provides a brief technical summary of the details behind data stores associated with the processes in Diagram 2.
Table 2. Technical details of the Data stores in the GDI-EE system visualised in Diagram 2.
| Data store | Data format | Contains | Encrypted | Associated process | Description |
|---|---|---|---|---|---|
| Dataset package | .tar.c4gh | Metadata, variant data, samples data (optional) | crypt4gh | Dataset package preparation | Prepared by data-provider and added in the GDI project bucket |
| Variant files | parquet | validated genomic variant data | crypt4gh | Dataset ingestion; Variant files validation; Data search | Uploaded to node S3 upon successful validation; a pre-requisite for dataset registration |
| Dataset manifest file | yaml | validated dataset metadata | crypt4gh | Dataset ingestion; dataset verification | Uploaded to node S3 upon successful validation; a pre-requisite for dataset registration |
| Samples file (optional) | parquet | validated samples phenotypic data | crypt4gh | Dataset ingestion; samples file validation (optional) | Uploaded to node S3 upon successful validation; a pre-requisite only for subject-level dataset registration |
| Database tables | database record | dataset title, description, internal ID, catalog, dataset files; samples count, individuals count, min/max age; record count | - | Dataset registration | Upon successful validation, Dataset is updated from UNREGISTERED to REGISTERED |
| Dataset status field | database record | dataset status | - | Dataset management; Catalogue data preparation; Data search | Possible statuses are: UNREGISTERED, REGISTERED, VISIBLE, HIDDEN, ARCHIVED; VISIBLE datasets’ properties will be published in a data catalogue at FAIR Data Point service |
| Access application | text document | research project proposal | - | Data access management | Managed according to national and institutional agreements; Remains to be specified at central level across Europe |
| Data cohort manifest | yaml | research cohort metadata | crypt4gh | Data transfer to SAPU | Prepared for dataset that has been granted access and permission for analysis in SAPU |
| Data cohort files | vcf, fastq, cram, bam, other | research cohort genomic data | crypt4gh | Data transfer to SAPU | Original genomic data files in data-provider storage that have a permission to be transferred to SAPU |
| Analysis results | analysed data (image, table, statistics) | results file | - | Data analysis in SAPU | Generated by the researcher during data analysis; stay in SAPU until permission for export is granted |
Data protection aspects
The data protection by design approach of GDI-EE system includes the following main features: 1) use of pseudonymised data where the de-pseudonymisation keys are stored outside GDI system, 2) transfer of data in an encrypted format where decryption keys are kept separately by secret management software, 3) using file checksum computation to protect and check for genomic data integrity within the system.
The most vulnerable component of the GDI architecture in terms of potential data leakage and privacy-risk is the Beacon data that must be exposed for requests via User Portal. However, the privacy-risk does not concern the queries performed via Allele Frequency Browser that is connected to aggregated Beacon which only delivers summary-level statistics (no individual data registered in the system). In fact, the first implementation of GDI is limited to queries of aggregated Genome of Europe data only, and restricting access to personal data within the national SPE or SAPU in our case. This is due to legal aspects that await decisions by the European Commission.