Dataset Preparation
- Version: 0.5
- Status: DRAFT
- Created: 2026-03-02
- Updated: 2026-03-13
- To be consulted:
- Kersti Jääger (kersti.jaager@ut.ee)
General process
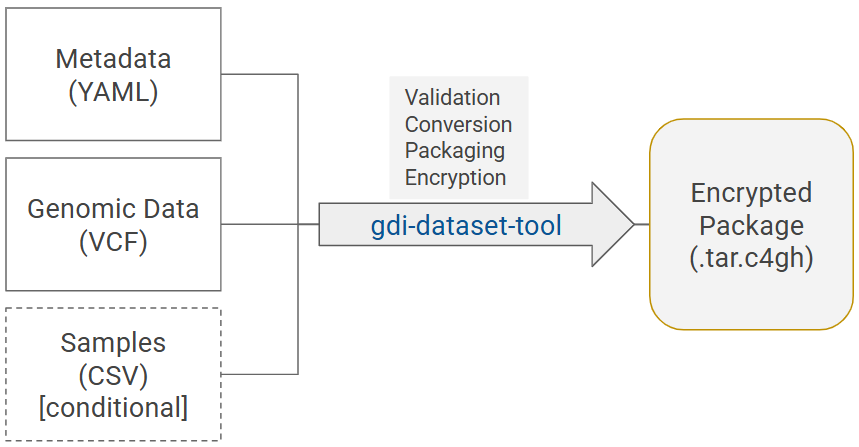
Diagram 1. General process of dataset preparation: gdi-dataset-tool validates, transforms and collects input data files - a metadata file, a VCF file, and a samples file (only for individual-level data) - into a dataset package, and encrypts the package for secure submission.
Preparing dataset files
Data-providers first need to generate the input files for the gdi-dataset-tool - a script that turns data files into a format compatible with GDI-EE. The tool accepts:
- VCF files as genomic data files
- Metadata file in YAML format describing the dataset (package.yaml)
- Sample file in a CSV file (only when sharing individual-level data)
1. What to consider when preparing a VCF file?
- The input VCF file can either be uncompressed (.vcf) or compressed (.vcf.gz).
- Chromosomes must be represented as 1–22, X, Y, M. Otherwise MT is turned into M and ‘chr’ prefix is removed.
- Only A/C/G/T/N are accepted for REF and ALT.
- VCF format/content (including INFO fields) must conform to a standard agreed for a project or use case to enable harmonisation and federation.
- For aggregate data: do not include genotype information in the VCF as all information for variants (ag allele-frequencies) is aggregated across samples/individuals.
Example VCF (aggregated)
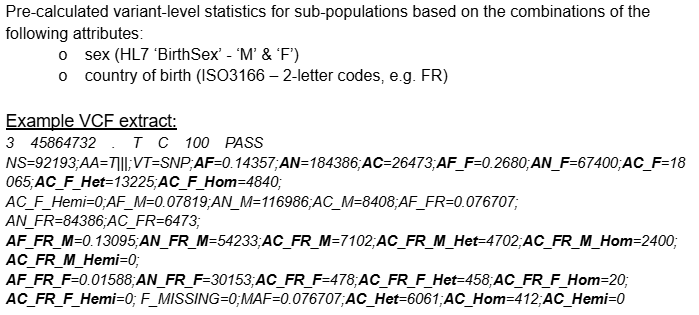

2. How to prepare the metadata file?
- Metadata describing the data needs to be in a package.yaml file.
- additionalFiles section in the package.yaml should be deleted if not used.
- Dataset metadata must specify whether a dataset includes aggregated or individual-level data (aggregated: true or false).
Metadata file (package.yaml)

3. When and what to include in the samples file?
- The samples file includes phenotypic information about the individuals.
- The file should be a .csv with the following delimiters allowed:
, ; \t |. - It should be provided together with a VCF that includes genotype information for these individuals (aggregated=false).
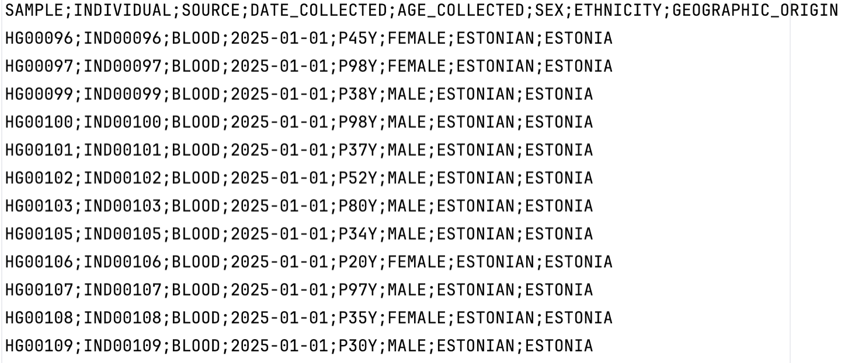
- The following column headers are required (order irrelevant): SAMPLE, INDIVIDUAL, SOURCE, DATE_COLLECTED, AGE_COLLECTED, SEX, ETHNICITY, GEOGRAPHIC_ORIGIN (final categories awaiting decision).
Samples file (csv)
Preparing encrypted dataset package
Once all the input files have been generated, gdi-dataset-tool can be run to generate a dataset package (Diagram 1) that can be submitted to GDI (to make it visible and findable in the User Portal).
The packaging involves:
- basic sanity checks for the content and match between the provided files (VCF, package.yaml, samples.csv)
- conversion of original files into Parquet files
- validating metadata to the manifest file
- packaging the files into a TAR file
- encryption of the TAR file using the public Crypt4GH key of the node and also the data-provider public key (which allows decryption of the package by the data-provider)
All these steps are carried out via a single command line function:
python3 -m gdi_dataset_tool package package_dir/package.yaml
The output file
The output file is an encrypted dataset package (<dataset.internalId>.tar.c4gh) that can be deposited in the Shared storage (Minio S3) or uploaded to GDI Estonia via local user portal. The package includes:
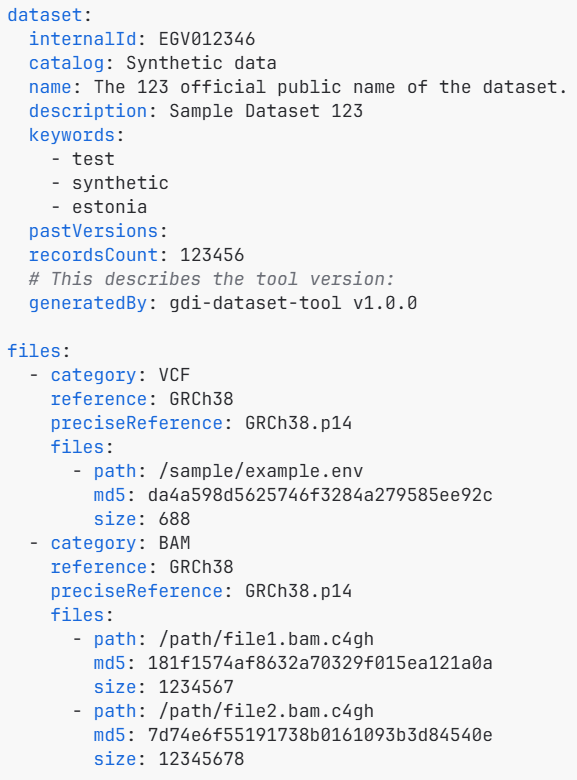
- A dataset manifest (manifest.yaml): Name, description, keywords; File paths, size, MD5-checksums
- Beacon data: variants-chr{c}.{p}.parquet or allele_freq-chr{c}.{p}.parquet
- Sample data (only for individual-level data): samples.parquet
- VCF-file headers (for internal use in the local portal): headers/{checksum}.vcf
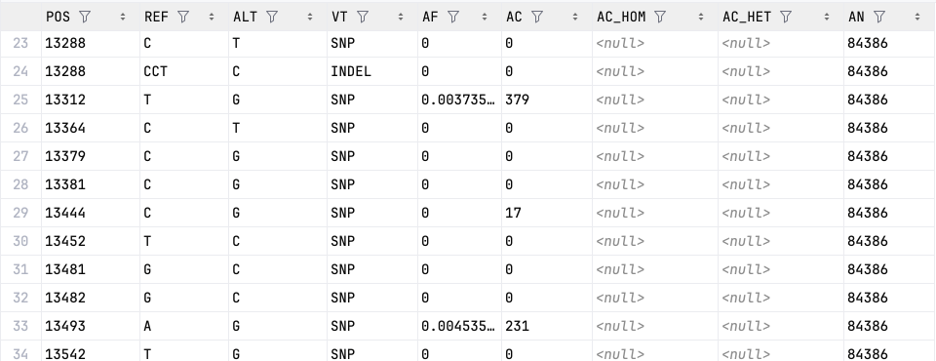
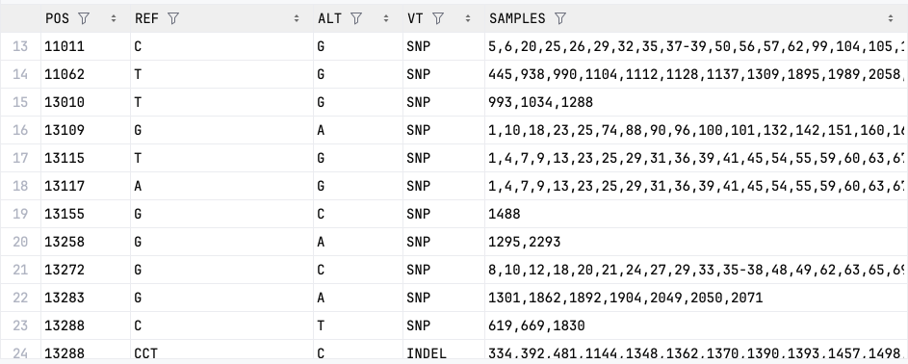
Examples of the Parquet files:
| allele_freq-chr{c}.{p}.parquet (aggregated data) | variants-chr{c}.{p}.parquet (individual-level data) |
|---|---|
 |
 |
The final manifest file:
See GDI-EE Technical quality checks for more details on validation and gdi-dataset-tool for detailed technical description of the tool.
Next steps
Dataset Submission to GDI Estonia